ThreaDomEx combines ThreaDom and DomEx into a unified on-line server system for more accurate and user-friendly domain predictions on sequences of both continuous and discontinuous domain structures.
In the search webpage, users are allowed to search the job by job ID or sequence. In addition, the navigation bar of all pages also provides a searching box which allows the user to search the job by the job ID.
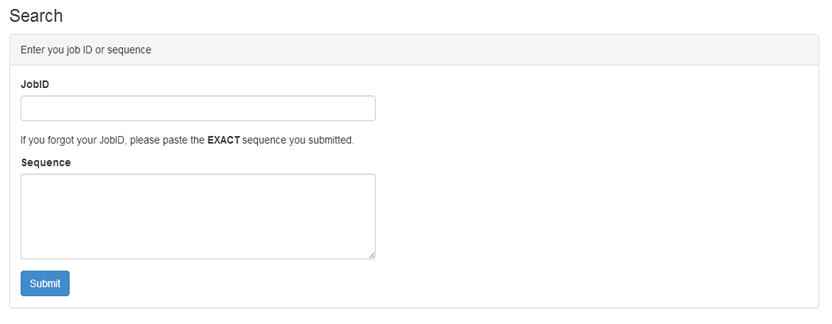
The ThreaDomEx takes the amino acid sequence in fasta format of the query protein as input. An optional email address to be informed about the job completion can be provided.

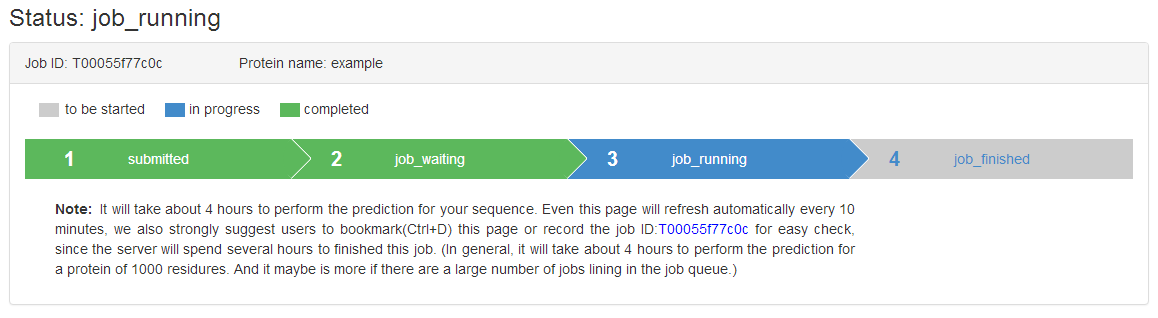
After submitting the job (by clicking the “Submit” button), a page containing a Job ID and a status bar that can be bookmarked will be displayed.
Output of the server consists of:
(a) the predicted domain boundaries
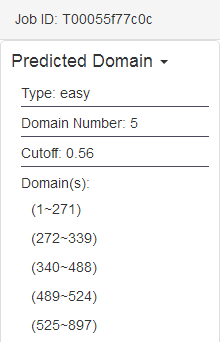

(b) the predicted discontinuous domains
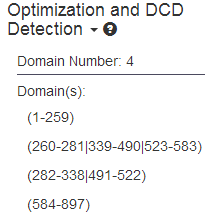

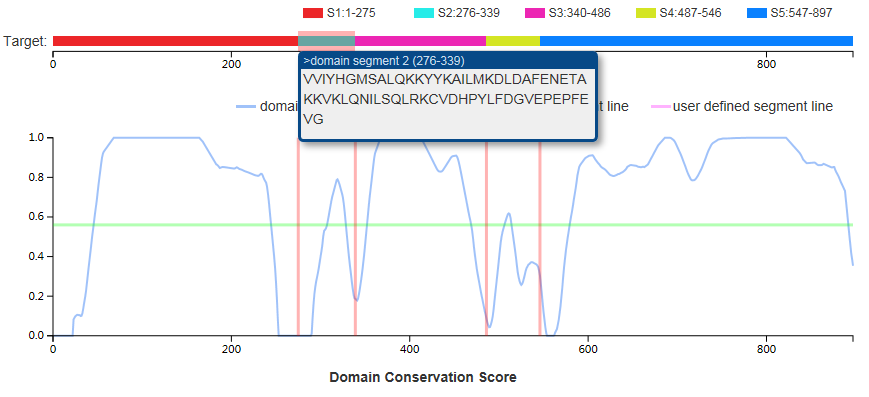
(c) the visualized distribution of domain conserve score

The whole sequence will be shown when hovering the mouse over the "Target" bar.

The segment sequence will be shown when hovering the mouse over the segment bar.
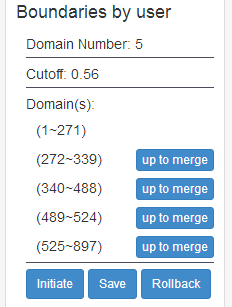
The server allows users to interactively edit, save, or re-detect the domain models of the proteins.
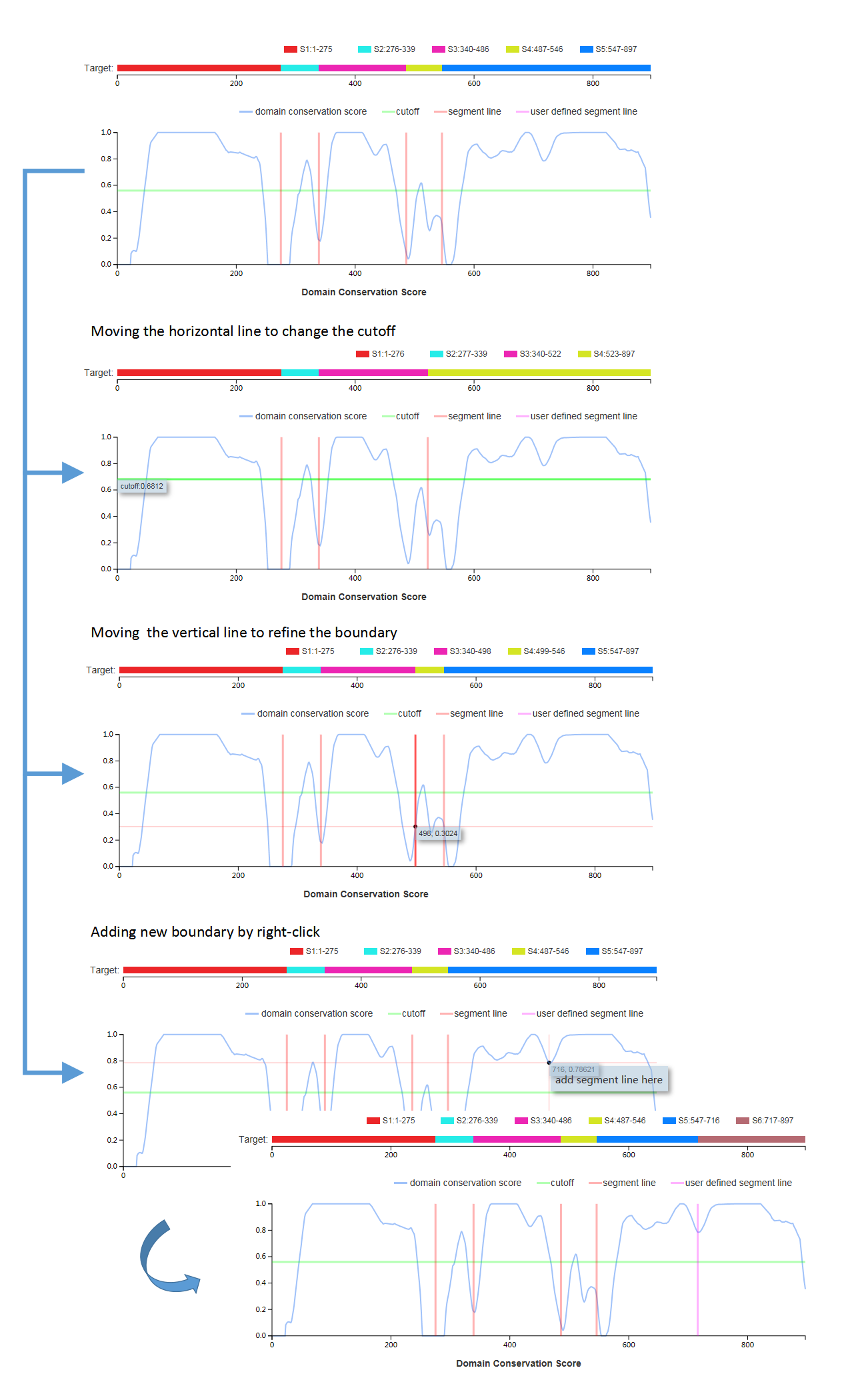
After the editing, the users can save the edit (by clicking the “Save” button) and redetect discontinuous domain(by clicking the “DCD Detect” button).
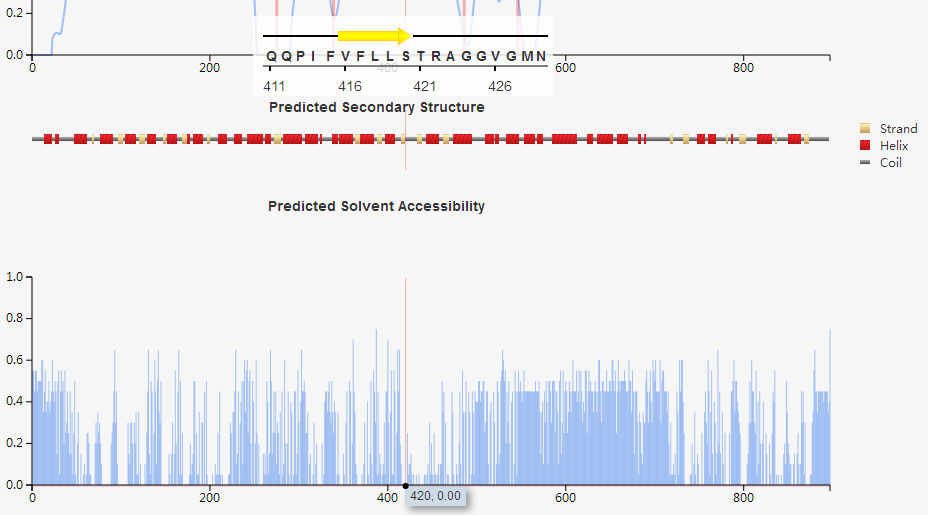
(d) predicted secondary structure and solvent accessibility
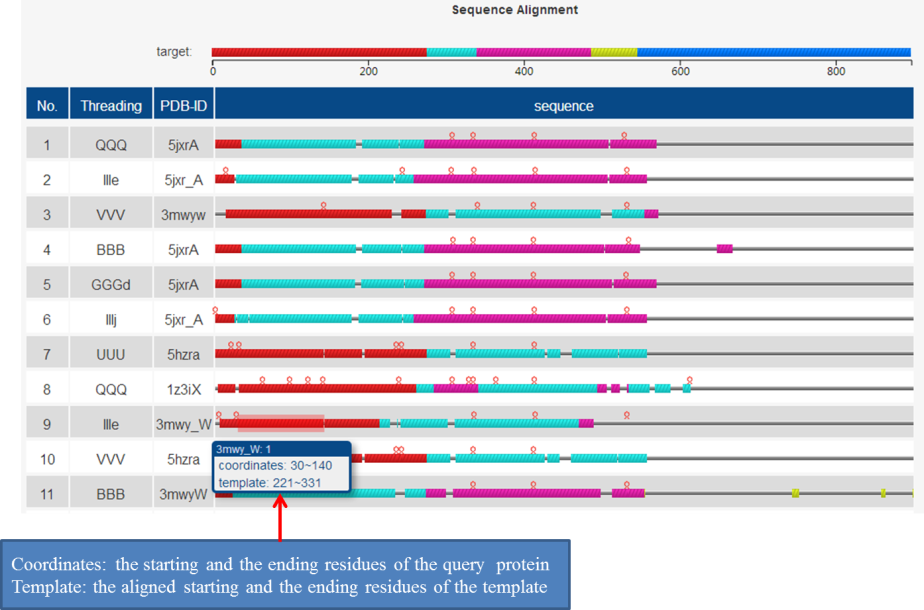
(e) the threading templates used by ThreaDomEx. Blocks with different color indicate different domains.